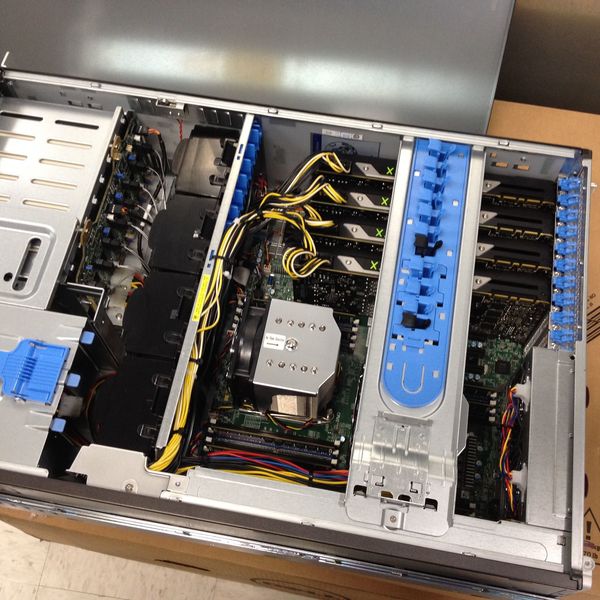
With cancer affecting millions of lives each year, Notre Dame scientists are working to develop personalized cancer vaccine therapies with the help of computational modeling. The recent acquisition of a General Purpose Graphics Processing Unit (GPGPU) computer cluster has significantly accelerated output for Notre Dame researchers. Led by Brian Baker, associate dean for research and graduate studies in the College of Science and professor of chemistry and biochemistry, an interdisciplinary team of biophysicists, biochemists and immunologists are using the GPGPU cluster to develop new immunotherapeutics. The cluster is maintained and housed by the Center for Research Computing at Union Station Technology Center, downtown South Bend.

Cory Ayres, a biochemistry graduate student who is co-advised by Baker and Steven Corcelli, associate professor of chemistry and biochemistry, leads the computational effort. Ayres focuses his research on molecular dynamics studies of T-Cell receptors (TCRs) and major histocompatibility complex proteins bound to peptide antigens (pMHCs). He explains that cells in the human body express proteins, and eventually, after several cellular processes, these proteins are broken up into multiple fragments called peptides. These peptides are then loaded onto an MHC protein and exported to the surface of the cell. Ayres further explained, “Once on the surface of the cell, T-Cells and their T-Cell receptor are able to sample the peptide presented by the MHC protein. If the peptide is recognized as non-self, the T-Cell will initiate an immune response against that cell. This overall process is a large component of what is termed adaptive immunity.”

Ayres uses the GPGPU cluster for molecular dynamics simulation in order to observe the motions of the TCR and pMHC proteins, and to determine the role that these motions play in molecular recognition. The new cluster provides 40 NVIDIA Titan X Cards in 10 Intel based servers. The primary software used for these simulations is the AMBER Molecular Dynamics Suite. “When I first started working on the project, I was working with CPUs only and could simulate about a nanosecond a day.” Ayres said. “Now I’m up to 1.2 microseconds a day, so I am seeing results at a factor of 1,000 times greater from when I started.” The scientists collaborated closely with the ND CRC, AMBER GPU Development Lead Dr. Ross Walker and the EXXACT Corporation.
Not alone in this endeavor, Baker and Ayres also collaborate with fellow researchers at Notre Dame, as well as the lab of Professor Pramod Srivastava from the Department of Immunology, University of Connecticut Health Center.
After sequencing cancer genomes, identifying the mutant proteins that are expressed, and predicting neoepitopes, Dr. Srivastava provides lists of candidate neoepitopes to the Notre Dame team. Ayres is then performs molecular dynamics simulations in order to help ascertain whether a given neoantigen will stimulate an anti-tumor immune response. Ultimately, this will help in identifying neoepitopes that may be useful as cancer vaccine components. In the near future, this work will be a component of a human clinical trial for ovarian cancer. “This cluster is really going to accelerate the throughput of our contribution to the project,” said Ayres. The ultimate goal for researchers is to develop patient specific, personalized vaccines. With the acquisition of this new cluster, their fight against cancer has been substantially accelerated.
For the full research paper, please visit The Journal of Experimental Medicine. The cluster was funded by a grant from the Carole and Ray Neag Comprehensive Cancer Center and Center for Immunotherapy of Cancer and Infectious Disease.
Originally published by at research.nd.edu on July 15, 2015.